Progress on Fluorescent Probes: Promising Biological and Clinical Applications
by Dr. Ozlem Dilek[[1]]
Husson University, 1 College Circle, Department of Biochemistry, Bangor, ME 04401, USA and Altinbas University, School of Medicine, Bakirkoy Istanbul 34147, Turkey.
Sponsored by LumiPedia
Reviewed by: Anonymous
Reviewed by: Anonymous
Date August 1st, 2019.
Contents
Fluorescent probes are molecules that absorb light at a specific wavelength and emit light of a different and longer wavelength (known as fluorescence), and are used to study and monitor biological systems in vitro and in vivo. Fluorescent molecules, usually called as fluorophores, are mainly used to investigate on a specific parameter that is physical, chemical or structural such as like pHor the concentration of a species.For example, some probes that are sensitive to pH can be used to measure pH. Some probes with long excitation and emission wavelengths can be performed in tissues or live cells, which display autofluorescence at short excitation wavelengths. Moreover, fluorophores that are sensitive to pH can be used to determine pH levels in biological systems. Fluorophores that have long excitation and emission wavelengths can be used in tissues, which show autofluorescence at short excitation wavelengths.
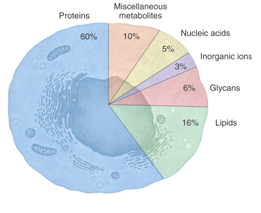
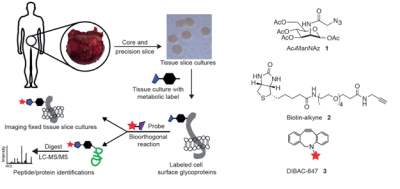
Thousands of fluorescent probes have been developed and known. Fluorophores have been designed and designed to probe specific particular site of a biomolecule to monitor the functionality, binding activity, structural behaviors, flexibility or conformational changes of a certain biomolecules inside cell network. This review contains an overview of the various types of fluorescent probes with their spectral properties, and applications. Fluorophores can be broadly divided into two main classes—intrinsic and extrinsic. Intrinsic fluorophores are those that naturally exist and are known as NADPH, FAD, FMN, pyridoxyl chlorophyll and aromatic amino acids (Tyrosine, Phenylalanine, tyrptophan), pteridines, and proteins. Proteins are the ones that are frequently studied natural fluorescence systems. [1-2] Intrinsic fluorescent pyridine nucleotides and flavoproteins are typically assigned as noninvasive optical fluroescent probes. Excitation region for NAD(P)H are at 365 nm and emit between 400-500 nm. However, flavoproteins excite at 450 nm and emit between 500-600 nm.[1-2] Fluorescent molecules, known as fluorophores, have light sensitive fluorescence spectral characteristics. Each fluorophore has distinct spectroscopic properties, which can be used to determine which fluorophore would be useful for a particular experimental or biological system. For example, some proteins or small molecules in cells (Figure 1)[3-4]are naturally fluorescent; this is called intrinsic fluorescence or autofluorescence [e.g., green fluorescent protein(GFP)]. Proteins, nucleic acids, lipids, or small molecules can be labeled with an extrinsic fluorophore—a fluorescent dye (or imaging agent)—which can be a small molecule, a protein or a quantum dot or a nanoparticle. [5] For example, Bertozzi and coworkers developed several chemical tools and strategies for labeling glycosylation in biological systems, so that these technologies will promote alternative strategies for diagnosing diseases and further treatments.[5-6](Figure 2) Recently, Bertozzi and coworkers reported a new biorthogonal strategy that was the first application of glycoproteomics to human tissues cultured ex vivo. [7] Nagano and coworkers reviewed and categorized the topic of fluorescent probes by focusing on focus on organic small molecule fluorophores and their use as labels and sensors for bioimaging. Probes using fluorescent proteins, quantum dots, lanthanide ions. [8]
1 Recent fluorescent probes and their applications[25]:
1.1 BODIPY dyes:
Boron-dipyrromethene (BODIPY) derivatives are known as one of the most popular family of fluorescent dyes in the field of bioimaging. One of their main advantages, e. g. large fluorescence quantum yield (QY), easy modification, high selectivity and specificity, sharp fluorescence emission, high extinction coefficient and excellent photostability[8-15].
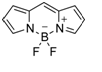
BODIPY originally synthesized 1968 by Triebs and Kreuzer [16] and developed as a laser dyeand used for many different applications including with biological labeling. Early applications of BODIPY compounds were covered in many publications. [17-20] BODIPY molecules (Figure 3) usually show narrow absorption bands with higher absorption maxima (500 nm or more).In special cases, where the BODIPY core is highly conjugated so that the absorption amd emission spectra move to the near infrared region. [22-24] Compared to regular very well-known fluorescent dyes such as fluorescein and rhodamine, borondipyrromethene (BODIPY) and boron-azabipyrromethene (aza-BODIPY) have unique features such as strong absorption in the visible region, narrow emission bands with high quantum yields, and excellent physiological stability. [61-65] The absorption spectra for BODIPY compounds are only slightly affected by solvent. For example, there is a small hypsochromic shift in the absorption maximum for 1 in changing solvent from toluene. Emission spectra of BODIPY dye is relatively reasonably matching with its absorption maxima and also display high fluorescence intensities compared to other dyes. The Stokes shift (SS) is shown to indicate only a modest change in structure following the S0 to S1 transition and vibrational relaxation. Some interesting clinical applications were performed with BODIPY derivatives such mitochondria-targeted fluorescent dyes mainly utilized delocalized lipophilic cations (DLC) as cargo groups in consideration of the negative mitochondrial membrane potential. [20-23] The most widely used mitochondria-targeted delivery moiety nowadays is cationic triphenylphosphine (TPP+). Tao et al., reported two novel mitochondria-targeted fluorescent probes, such as TPP-BODIPY and TEA-BODIPY and they used the cargo TPP+ and TEA+ with the fluorophore BODIPY by making a flexible carbon chain, respectively. The cationic groups can therefore guide BODIPY into mitochondria without losing its good spectroscopic and photochemical properties.
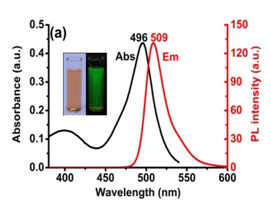
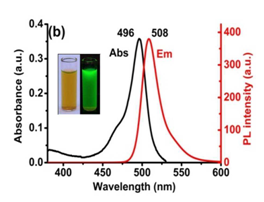
Talita et.al., recently published the design and characterization of fluorescent quinone-based BODIPY hybrid structures and showed their cytotoxicity against cancer and normal cell lines and also performed some mechanistic studies by lipid peroxidation.[24] They additionally determined the reduced (GSH) and oxidized (GSSG) glutathione and imaged their subcellular localization by confocal microscopy. They studied two fluorescent BODIPY based probes, namely TPP-BODIPY and TEA-BODIPY, which were applied to mitochondria-targeted imaging in living cells. They possessed many advantages, including superior brightness (Figure 4), high photostability, low cytotoxicity and fast internalization, thus they are one of the alternative probes, which are good candidates for mitochondria-specific imaging.
Koleman et al., also mentionedthe current advances in BODIPY chemistry also allow the synthesis of red-shifted BODIPYs [8] such as the advantages of far-red and near-IR probes for biological applications due to their long range absorption and emission wavelength and low autofluorescence, minimal phototoxicity. Furthermore, red-shifted probes can also be suitable for in vivo imaging, which is highly useful for practical applications due to deeper tissue penetration of the incoming and outgoing light. The major drawback of BODIPY derivatives is their high hydrophobicity leading to low water solubility. For example, various hydrophilic groups, such as sulfonate, pyridinium, glycol, and carboxylate, were appended to the core of dyes to increase their aqueous solubility. [53-55] Introduction of a functional group such as the ester or carboxylic acid into dyes could increase their membrane permeability. [56-57] However, this problem can be simply addressed by decorating the core structure with hydrophilic moieties through well- established examples of BODIPY chemistry.[59-61, 66] Wang et al., mentioned this solubility issue on his work and studied Boron-dipyrromethene (BODIPY) as a model fluorescent dye, for the first time, they designed a new strategy for the rapid production of a water-soluble and membrane-permeable dye by mixing with an amphiphilic protein named hydrophobin. Their data exhibit BODIPY could be effectively solubilized and dispersed in 200 μg/mL hydrophobin by simple mixing and sonication. [58] There are several excellent reviews on literature regarding the chemistry and spectroscopic properties of BODIPYs as well as some others highlighting BODIPY-based fluorescent probes.[14-18]
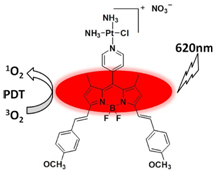
Alternatively, in a different approach, Liu and his co-workers synthesized a NIR fluorescent BODIPY-Platinum conjugates (BODIPY-Pt) (Figure 5), which eventually produces NIR fluorescence, chemotherapy and photodynamic effect. Usage of platinum significantly increased the photodynamic effect of BODIPY. They have also monitored cellular uptake and distribution of the BODIPY-Pt by confocal laser scanning microscopy(CLSM). In addition, they further studied its phototoxicity and cytotoxicity against human cervical carcinoma (HeLa) and human hepatocellular liver carcinoma (HepG2) cell line. This approach, thereforeshows a good example of the distribution of the BODIPY-Pt in tumor tissue according to NIR fluorescence imaging in vivo.[14]
1.2 Coumarin dyes:
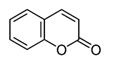
The coumarin scaffold (Figure 6) is a very well-known structure in organic chemistry, taking place in various natural products and pharmacological agents. [31] Substitution at the 7-position with electron-donating groups yields highly fluorescent molecules. The prototypical coumarin fluorophore is 4-methyl-7-hydroxycoumarin, which has an absorption maximum (λ[math]_{abs}[/math] = 360 nm, ε = 1.7 × 10[math]^{4}[/math] M[math]^{−1}[/math]cm[math]^{−1}[/math]) and emitting a blue light (λ[math]_{em}[/math] = 450 nm, Φ = 0.63).All these relevant colorful agents are the basis for many fluorescent labels, fluorescent sensors, and fluorogenic enzyme substrates. One of the most popular chemical modifications on coumarin core is halogenation, which significantly decreases the pKa of the phenolic hydroxyl groups and thus limits pH-sensitivity. [32]
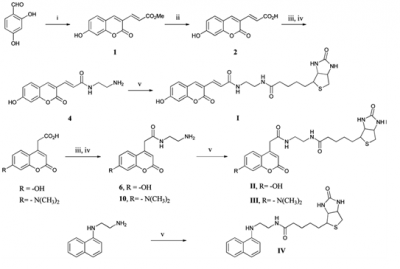
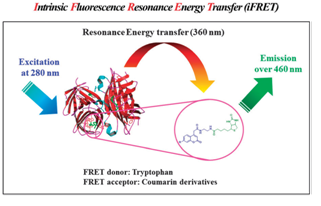
Coumarin scaffolds have been extensively used in the preparation of fluorescent dye-protein conjugates and as the basis of fluorogenic enzyme substrates. Due to their highly demanding usage in biological assays, 7-hydroxycoumarins and their conjugates are not fully deprotonated-not maximally fluorescentexcept the dye stays in basic environment having a pHof 9 or higher. Moreover, many protein conjugates of 7-hydroxycoumarin are not only unstable, but also needs basic conditions to reach maximal fluorescence intensity of the dye. [33] Coumarin based dyes are well-explored and various reports has been published with their synthesis and their applications. [40-41] The effects of substituents at the 3- and 7-position of coumarin drastically change the fluorescence characteristics by changing the lowest excited states of the fluorophore. Kim and co-authors synthesized three coumarin derivatives, which had electron-donating groups at the 7-position and 3- or 4-position. They actually used them for biotin conjugation. The synthesis of the coumarin derivatives was prepared either via Pechmann- or Knoevenagel condensation. Conjugation of fluorescent coumarin scaffolds with biotin via EDC mediated amide bond formation, to yield fluorescent probes I, II, III and IV (Scheme 1).Coumarin based fluorophores in this study were used with iFRET technique which represents a label-free and real-time imaging method to study biological systems. Three fluorescent probes were prepared and conjugated to biotin. The biotinylated fluorescent probes were used in iFRETstudies with purified streptavidin. Experimental data confirmed that the obtained probes are effective iFRET acceptors for tryptophan-containing proteins. Coumarin-based probe II was identified as a potent iFRET acceptor (Figure 7) for tryptophan containing proteins. [39]
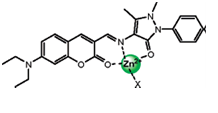
Different aspects of fluorescence mechanism between recognitionand signal response based units is also a developing area to design new fluorescent probes or chemosensors. Indeed, until now, a number of signaling mechanisms have been developed and widely applied for the optical detection of biological species in live cells or in vitro. Wu et al., designed a coumarin fluorescent dye, coumarin and the antipyrine derivative acts as an additional chelating moiety on the chromophore system. Both chemical sides are linked by a C=N bond to form a potential fluorescent-sensing molecule (L) for metal cations. [42] They synthesized the Compound L according to the new approach as shown in Figure 8 and monitored the turn off/on fluorescence mechanism—chelation of metal through coumarin moiety.
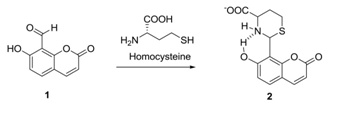
Yap et al. reportedthe possibility of a chemoassay using a coumarin-based fluorescent probe (Scheme 2) to quantify homocysteine in the Parkinson Disease patients’ blood serum. They have designed a rapid monitoring of homocysteine, methylmalonic acid, and vitamin B12 using LCMS. Quantification of CNS (central nervous system) disease biomarker was determined in the blood serum of Parkinson’s disease patients with and without the levodopa treatment and compared against the healthy data. [43]
Detection of oxidative stress in cells has also been one of the major clinical subjects to investigate. [44] Oxidative stress(OS)is characterized by accumulation of reactive oxygen species (ROS) in the cellular systems. [45] Increased ROSlevels can cause some damages in biomolecules including proteins, lipids, nucleicacids and carbohydrates. [46] Detection of acuteorchronicpatho physiological conditions, such as Parkinson’s and Alzheimer’s diseases, atherosclerosis, heart failure, end othelial dysfunction, and inflammatory disorders are the fundamental clinical cases to be diagnosed with possible imaging techniques. Vemula et al., designed a coumarin hydrazide-based fluorescence technique which can analyze proteincarbonylation in cells. [44]
The carbonylated species in fixed cells can be detected by fluorescence microscopy. DNPH (dinitrophenylhydrazine) is frequently used to visualize cellular systems by immunochemical analysis with anti-DNP antibodies. [47] Mukherjeeet al.,reported an article on detection of carbonylation-oxidative stress on live mammalian cells with a hydrazone conjugated system. She and co-workers have used a hydrazine-functionalized fluorophore has been to observe oxidative stress induced carbonylation of lipids and proteins in live mammalian cells. Coumarin hydrazine based fluorophore has good spectroscopic properties to detect cellular carbonyls in a single step procedure using confocal microscopy. [48]
Coumarins are also used as effective fluorophores to detect bithiols in living cells due to their high fluorescence quantum yield, good water solubility, low toxicity, and better cell permeability. [49-51]Li and his co-workers synthesized two fluorescent probes (PCS and NCS) for biothiol detection in aqueous solution based on the mechanism of intramolecular charge-transfer (ICT). [52]
1.3 NIR dyes:
Compared to other long wavelength fluorescent dyes, NIR dyes are getting much more attention due to their applicability (transperancy of tissues in the wavelength region) in in vivo imaging in animals and humans (Figure 9). For example, fluorescein derivatives in which the carboxyl group is replaced with other functional groups, termed TokyoGreens (TGs), were not created until the twenty-first century. Bright and photostable dyes with red to near-infrared (NIR) fluorescence are especially mostly modified to address the membrane permeability. From both biophysical and cell biology points of view, deep tissue penetration of NIR light have some advantageous over UV–vis light such as it does not exhibit any significant interference with light from biomolecules when monitoring in vivo. Weissleder and his coworkers compared NIR light emission with that of green and red ones in mice and found that NIR is a superior light in terms of background autofluorescencein the animal model. [67] NIR light characteristics show that these NIR fluorophores with a non-invasive imaging tool can be very useful in vivo. Due to these imaging limitations, many reviews of NIR probes for bioimaging have been reported recently. [67-73] Chin et al. studied the molecular structures of NIR fluorophores and then describe their enzymatic reactions with peptidases from the viewpoint of chemical design and reaction mechanism.[74]
BODIPY molecules can be designed with particular unique properties such as Stokes shift, high fluorescence quantum yield that is independent of the environment, sharp excitation and emission peaks, and high solubility in many organic solvents. The combination of these characteristics makes BODIPY a suitable candidate for imaging applications. Akkaya et al. showed the synthesis of Near-IR fluorescent BODIPY derivatives by introducing conjugated electron-rich aryl groups at 3,5-positions of the BODIPY scaffold, which could have longer wavelength emission range than previously reported BODIPYs. [76-77]
1.4 Squaraine-based probes:
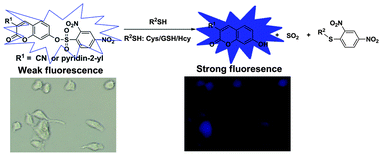
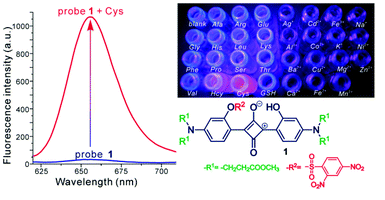
Squaraines are dyes with an electron deficient four-membered ring and a resonance stabilized zwitterionic structure. The central ring is typically appended with donor moieties to afford a donor–acceptor–donor motif.D–π–A–π–D conjugation structure core on squaraine makes the fluorophore distinguished more than the other known NIR fluorophores because of their better photophysical properties such as high extinction coefficient, high quantum yield, high photostability and long emission wavelength in the red or near-infrared region. It was reported that the complete fluorescence quenching was observed in the presence of ten equivalents of cysteine. [80] Liu and coworkers reported a series of squaraines with 2,4-dinitrobenzenesulfonyl group on its structure in order to improve solubility and lower the response time for imaging biothiol probes. (Figure 10) These molecules exhibit fluorescence at red emission spectral region. Furthermore, membrane permeability issue was overcome by this synthetic probe to detect fluorescence for thiol species in living cells so that this probe was selected as a good candidate for biothiol detection process in living cells imaging. [80]
Suzuki and co-workers introduced very effective water-solubilizing sulfonate moieties into asquaraine framework. [80]it was reported that squarainetetrasulfonate fluorophore on a bovine serum albumin (BSA) was analyzed as conjugate which exhibited absorption maxima at 787 nm and emission maxima at 812 nm with a quantum yield of 0.08 inaqueous solvent. The authors concluded that synthesized dyes can be usefulin protein detection as covalent labeling probe and can also be applicable as contrastagents for in vivo imaging. Klymchenkoet al. reported different group of fluorogenic dyes based on asquaraine dimer, which unfolded in the organic environment from the aqueous folded structure so that the fluorescence turn on was initially detected in the hydrophobic environment. [79]
1.5 Cyanine-based probes:
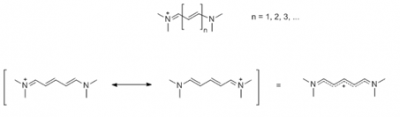
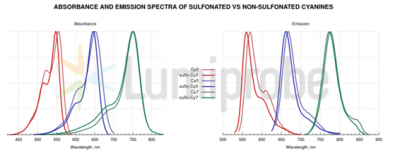
Near-infrared fluorescence (NIRF) imaging agents have high extinction coefficients, large Stokes’ shifts, and produce strong fluorescence emission at the range of 700 to 1000 nm[81], used formostly in vivo cancer diagnosis. Strong ligand labeling, high signal strength, less background fluorescence and tissue absorbance are the main advantages compared to other fluorophores. [82-84] Cyanine dyes (Figure 11) was introduced the first time in 1857 by C. G. Williams. [85] Several NIRF dyes have become commercially available, such as Cy5.5[86]and IRDye800-CW[39]which have been coupled with peptides or antibodies. These couplings frequently used for site-targeted monitoring of neoplastic cancers in animal models. Indocyanine green (ICG), the only NIRF agent approved by the FDA in the United States for medical diagnostic application, has been widely used to differentiate benign from the malignant diseases in the clinic. [88] Sulfonated and non-sulfonated cyanines (Figure 9) exhibit very similar fluorescent properties. However, it should be noted that non-sulfonated cyanines must be dissolved in an organic co-solvent (DMF or DMSO) before use. Sulfo-Cyanine reagents can be used in purely aqueous conditions. They are differentiated in terms of purification methods: when dialysis against water or aqueous buffer is used for purificationa and sulfo-Cyanine must be used to remove of unreacted excess. Both sulfo- and non-sulfo cyanine based reactions can be purified by gel filtration, chromatography (HPLC, FPLC, ion exchange), or electrophoresis.[97]
Conventional NIRF dyes tend to be used with chemical conjugation of cancer-specific targeting ligands, such as metabolic substrates, cell-surface peptides, growth factors, antibodies, and cancer-specific cell-surface biomarkers for cancer imaging in vivo models. [90-94]Thus, only some certain cancers can be identified by these imaging probes due to tumor cell heterogeneity. Additionally, site-specific imaging or selective-targeting of tumor-targeted ligands may be altered withdifferent chemical conjugation systems. [95]
Shi et al., reported a review on NIRF imaging agents (cyanine dyes for tumor imaging, photodynamic therapy. [96] However, most of these dyes exhibit fast degradation or decomposition with low solubility in polar solution, low quantum yield and tumor-targeting specificity issues. [89]
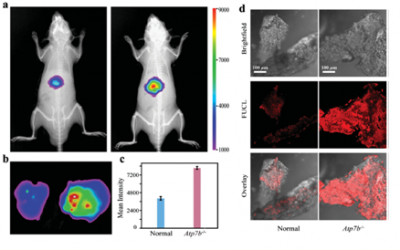
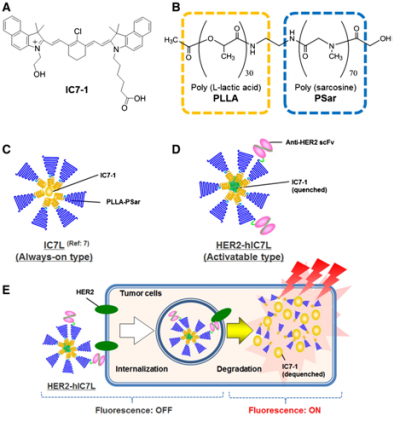
Shimuzu and his co-workers also first prepared lactosomes encapsulating IC7-1 with different concentrations, and evaluated both their quenching and dequenching properties in in vitro experiments. (Figure 12) They developed HER2-hIC7L that shows self-quenched fluorescence and specifically interacts with HER2 expressed on tumor cells, followed by the internalization in these cells. These results show that an antibody-conjugated lactosome that encapsulates higher concentrated IC7-1 would be a useful NIR probe that is applicable for use in noninvasive in vivo optical imaging for specific detection of target biomolecules expressed in tumors. [98] (Figure 13)
1.6 Metal-based probes:
Several probes have also been designed for fluorescence signaling and widely used for the detection of biological systems including metal ions due to their high sensitivity and low cost and suitability for in vivo imaging. [99-113] Wu et al., reported a review perspective on how fluorescent probe-metal ions interactions have been developed in recent years. [114](Figure 14) Their photophysicalproperties depends on fluorescence mechanism changes such as intramolecular charge transfer (ICT), photoinduced electrontransfer (PET), Förster resonance energy transfer (FRET)[[2]], excited-state intramolecular proton transfer (ESIPT) and chelation induced enhanced fluorescence (CHEF), have been widely used for probe design to be used in biological imaging. Alkali metal ions include lithium (Li[math]^{+}[/math]), sodium (Na[math]^{+}[/math]), potassium (K[math]^{+}[/math]), rubidium (Rb[math]^{+}[/math]), cesium (Cs[math]^{+}[/math]) and francium (Fr[math]^{+}[/math]) ions. Francium, which is highly radioactive and toxic and it has no biological role. [115-116] Cesium is not an essential element for human beings, but it is very well-known in biochemistry and physiology. Most of the other alkali metal ions play important roles in pathophysiological systems. Lithium salts are essentially used in medical and clinical applications such as in bipolar disorders and manic-depressive psychosis cases. Sodium and potassiumare very important critical ions in living organismsbecause of their roles in many biological processes. Furthermore, imbalances of sodium and potassium are closelyrelated to many diseases such as Alzheimer’s disease (AD),Wilson’s disease (Figure 15), anorexia, heart disease and diabetes.[117-120]
Crown ethers are one of the largest probe categories, which are used with alkali metals to monitor biological processes. However, the NIR fluorescence probes for alkali metal ions are very less known and needed for future biological applications.
2 Fluorescence Imaging Techniques in Drug Delivery and Cancer Therapy:
Etrych and his coworkers recently published an article on applications of different fluorescence imaging techniques in the field of drug delivery and anticancer therapy. They mostly focused on the potential applications and limitations of non-invasive imaging techniques. [34] In these studies, in vivo FI is typically based on the illumination of a target tissue with a light source of a specific wavelength or wavelength range (e.g., from ultraviolet to infrared) that is able to excite fluorophores. The excitation light has to penetrate from membranesto several tissue layers to reach with the possible suitable fluorophores. The light, is therefore, can be reflected or scattered at some certain points. The main advantages of FI are easy set-up, less ionizing radiation, long-term monitoring ability with the use of fluorophores. [35-36]
In addition to monitoring them in anticancer drug delivery and tumor biology research, understanding the malignancies or other neoplasms by using photodynamic therapy (PDT)in combination with DDSs is a very well-known treatment. [37] In this technique, A nontoxic photosensitizer-DDS conjugate accumulates at a target site after administration. They irradiate the target tumor with the appropriate light which activates the photosensitizer and, immediately, induces the generation of toxic substances such as known as reactive oxygen species. These molecules often trigger cell death and cause to the therapeutic treatment of disease. For example, porphyrin-based nanoparticles can act as a singlet oxygen converter. Additionally, they are mostly used in photothermal therapy (PTT)and the light is transferred into destructive heat inside tumor tissue. [38]In short, the specific properties of each fluorophore determines a wide range of in vivo theranostics. Fluorescence-based techniques have been frequently used in clinical practice in surgery, phototherapy, and the detection of inflammatory events.[37] Liu et al., worked on hemicyanine NIR probe 41 which also shows a selective fluorescence turn in response to Cu2+ and applied to detecting Cu2+ in living Siha cells.[114]
Moreover, to image physiological and pathological aspects of diseases at a level of different stages, metal ions might be essential to overcome some problems such as turn-on targeting. However, there are still unknown problems and challenges exist to image certain functionalities at a specific site of biological systems. For instance, no NIR fluorescence probe with high selectivity and sensitivity for some metals has been designed and applied successfully for biological systems thus far.
Commonly used NIR fluorophores such as BODIPYs have low stabilities and solubility. Thus, a great demand is needed for the development of new NIR fluorophores that do not have these disadvantages. Some NIR probes are not effective enough to detect metal ions in vivo. Also, NIR probes possess some advantages such as applicability in lower energies, higher solubility in water, less fluorescence background after binding. However, all these advantages need to be reflected in in vivo experiments. Therefore, the development of water-soluble, stable NIR fluorescence probes would be very important in medicine and biological fields.
To analyze the exact mutations on specific sites on cancer cells is very challenging. Cancer therapies target not only the malignant cells but also the healthy cells. Most of the well-known anticancer agents cause some side effects on patients. Researchers and clinicians work to discover biomarkers and molecular tools that can identify cancers at a certain level. Addition to these therapies, there is a significant need to develop new tools that will allow monitoring cancer subtypes in an accurate and rapid ways. Bogyo and his coworkers summarized the new developments on molecular agents, which are recently used in clinical applications in detail. [122] Researchers pointed out that many of the available chemical probes have the potential to become clinically available in the next 5–10 years if they are potentially applicable in in vivo studies. By using the suitable contrast agents or small fluorescent probes, surgical procedures will proceed faster and significantly will also have a huge impact in the success of pre-clinical and post-clinical processes. Rodriguez et al., also reported a broad review of the current state of fluorescent probe development and applications in mammalian systems and discussed the rapid adoption of optogenetic approaches and new advances towards manipulation of biological pathways or monitoring the intracellular events by fluorescence. [123] Urano and his colleagues also reviewed the recent trends on tumor-targeted fluorescence imaging probes and their clinical/preclinical applications. They also reported their recent work on activatable fluorescence probes for tumors expressing aminopeptidases and glycosidases. Their results indicate that probes enable rapid and sensitive detection of tiny tumors smaller than 1 mm in diameter, both in vivo and in clinical samples. [124]
The design of new fluorescent probes targeted to tumor-specific molecules could provide a significant improvements in surgical and clinical procesures and will enhance the life quality of the patients in the long term.Having a powerful new fluorescent imaging technique to detect and analyze the cancerous lesions will overcome the problems resulting from current long and painful procedures. In the past few years, various types of new fluorescence probes for site-specific labeling or imaging of tumor imaging have been developed and this would open a new insight on imaging field such as combination of NIRF imaging with nuclear imaging techniques, such as PET and SPECT. The combination of these powerful imaging systems with creative fluorescent probes would additionally provide clinicians to image cancer-specific molecular targets with high resolution and sensitivity.
3 References:
1. Terai, T., Nagano T., CurrOpinChemBiol.2008 Oct;12(5):515-21.
2. Ellie Kohen, Joseph G. Hirschberg., Analytical Use of Fluorescent Probes in Oncology, 2012.
3. Alberts, B. Molecular Biology of the Cell (Garland Science, New York).
4. Apweiler, R., Hermjakob, H. & Sharon, N. Biochim. Biophys. Acta. 19991473, 4–8.
5. Prescher, JA.,Bertozzi CR. Nat Chem Biol.2005 Jun;1(1):13-21.
6. Sletten, E.M.; Bertozzi, C.R. Angew. Chem. Int. Ed. 2009, 48, 6974–6998.
7. Spiciarich, D. R.; Nolley, R.; Maund, S. L.; Purcell, S. C.; Herschel, J.; Iavarone, A. T.; Peehl, D. M.; Bertozzi, C. R.Angew. Chem. Int. Ed. Engl.2017,56,8992-8997.
8. Kolemen S., Akkaya U.E. Coordination Chemistry Reviews. July 2017
9. G. Ulrich, R. Ziessel, A. Harriman,Angew. Chem. Int. Ed. 47, 2008, 1184.
10. T. Kowada, H. Maeda, K. Kikuchi, Chem. Soc. Rev. 44, 2015, 4953.
11. Y. Ni, J. Wu, Org. Biomol. Chem. 12, 2014, 3774.
12. A. Loudet, K. Burgess. Chem. Rev. 107, 2007, 4891.
13. H. Lu, J. Mack, Y. Yang, Z. Shen.Chem. Soc. Rev. 2014, 43, 4778.
14. Liu Y, Li Z, Chen L, Xie Z., Dyes and Pigments, 2017, doi: 10.1016/j.dyepig.2017.01.075.
15. Benneston C. A., Copley G. Phys. Chem. Chem. Phys.2009, 11, 4124-4131.
16. A. Treibs and F.-H. Kreuzer, Justus Liebigs Ann. Chem.,1968, 718, 208.
17. T. L. Arbeloa, F. L. Arbeloa, I. L. Arbeloa, I. Garcia-Moreno,A. Costela, R. Sastre and F. Amat-Guerri, Chem. Phys. Lett.,1999, 299, 315.
18. G. Ulrich, R. Ziessel and A. Harriman, Angew. Chem., Int. Ed.,2008, 47, 1184; R. Ziessel, G. Ulrich and A. Harriman, New J. Chem., 2007, 31, 496.
19. K. Rurack,M. Kollmannsberger and J. Daub, New J. Chem.,2001,25, 289;W. Zhao and E.M. Carreira, Angew. Chem., Int. Ed., 2005,44, 1677; Z. Shen, H. Rohr, K. Rurack, H. Uno, M. Spieles, B. Shulz, G. Reck and N. Ono, Chem.–Eur. J.,2004, 10, 4853; W. Zhao and E. M. Carreira, Chem.–Eur. J.,2006, 12, 7254.
20. Terai, T.; Nagano, T. Pflugers Arch - Eur J Physiol. 2013, 465:347–359.
21. A. T. Hoye, J. E. Davoren, P. Wipf. Acc. Chem. Res.,2008, 41, pp. 87-97.
22. L. F. Yousif, K. M. Stewart, S. O. Kelley. ChemBioChem.,2009, 10, pp. 1939-1950.
23. Gao T, He H, Huang R, Zheng M, Wang F-F, Hu Y-J, Jiang F-L, Liu Y. Dyes and Pigments. 2017, doi: 10.1016/j.dyepig.2017.03.009.
24. Talita B. Gontijo, Rossimiriam P. de Freitas, Flavio S. Emery, Leandro F. Pedrosa, José B. Vieira Neto, Bruno C. Cavalcanti, Claudia Pessoa, Aaron King, Fabio de Moliner, Marc Vendrell, Eufrânio N. da Silva Júnior Bioorganic & Medicinal Chemistry Letters(2017-08-04), DOI:10.1016/j.bmcl.2017.08.007.
25. Lavis, D.L., T.R. Raines.ACS Chem. Biol., 2014, 9 (4), pp 855–866
26. FeiNiu, Xiu-Yun Song, Jin-Feng Hu, Wei Zuo, Ling-Lei Kong, Xiao-Feng Wang, Ning Han, Nai-Hong Chen, IMM-H004, Journal of Stroke and Cerebrovascular Diseases, Volume 26, Issue 10, 2017, Pages 2065-2073.
27. Xiaoqi Li, Fangjun Huo, Yongkang Yue, Yongbin Zhang, Caixia Yin. Sensors and Actuators B: Chemical, Volume 253, 2017, Pages 42-49
28. Clemens Zwergel, Brigitte Czepukojc, Emilie Evain-Bana, Zhanjie Xu, Giulia Stazi, Mattia Mori, Alexandros Patsilinakos, Antonello Mai, Bruno Botta, Rino Ragno, Denise Bagrel, Gilbert Kirsch, Peter Meiser, Claus Jacob, Mathias Montenarh, Sergio Valente.European Journal of Medicinal Chemistry, Volume 134, 2017, Pages 316-333.
29. Ye Li, Tongkai Chen, Xiaoqing Miao, Xiang Yi, Xueqing Wang, Haitao Zhao, Simon Ming-Yuen Lee, Ying Zheng. Pharmacological Research, Volume 125, Part B, 2017, Pages 246-257.
30. Bowen Li, Peilian Liu, Hao Wu, Xin Xie, Zelin Chen, Fang Zeng, Shuizhu Wu. Biomaterials, Volume 138, 2017, Pages 57-68.
31. Xiuli Yue, Qiang Zhang, Zhifei Dai.Advanced Drug Delivery Reviews, Volume 115, 2017, Pages 155-170.
32. Riveiro, M., De Kimpe, N., Moglioni, A., Vazquez, R., Monczor, F., Shayo, C., and Davio, C. Curr. Med. Chem. 2010, 17, 1325−1338.
33. Sun, W.-C., Gee, K. R., and Haugland, R. P. Bioorg. Med. Chem. Lett.1998, 8, 3107−3110.
34. Etrych, T., Lucas, H., Olga Janouskova, O., Chytil, P., Mueller, T., Mader, K. Journal of Controlled Release, Feb 2016.
35. J.C. Hebden, S.R. Arridge, D.T. Delpy. Phys Med Biol. 1997, 42, 825-840.
36. U. Mahmood, R. Weissleder. Mol. Cancer Ther. 2003, 2, 489-496.
37. “Photonics for Health Care", in Handbook of Biophotonics, Vol. 2, J. Popp, V.V.Tuchin, A. Chiou, and S.H. Heinemann, Eds., Wiley-VCH Weinheim, 2011.
38. Y.P. Li, T.Y. Lin, Y. Luo, Q.Q. Liu, W.W. Xiao, W.C. Guo, D. Lac, H.Y. Zhang, C.H., Feng, S. Wachsmann-Hogiu, J.H. Walton, S.R. Cherry, D.J. Rowland, D. Kukis, C.X. Pan, K.S. Lam. Nat Commun, 5, 2014.
39. Kim, J.H., Sumranjit, J., Kang, H.J., Chung, S.J. Mol Biosyst.2014 Jan;10(1):30-3.
40. H. Hussain, J. Hussain, A. Al-Harrasi and K. Krohn, Tetrahedron, 2012, 68(12), 2553–2578.
41. M. E. Riveiro, N. De Kimpe, A. Moglioni, R. Vazquez, F. Monczor, C. Shayo and C. Davio, Curr. Med. Chem, 2012, 17(13), 1325–1338.
42. Wu, J.S., Liu,WM., Fang Wang, XQ., Wang, PF., Tao,SL., Zhang, XH.,†Wu, SK., Lee, ST.Org. Lett., 2007, 9 (1), pp 33–36.
43. Yap, AC., Mahamad, UA., Lim,SY., Kim, HJ., Choo, YM. Sensors (Basel),2014 Nov; 14(11): 21140–21150.
44. Vemula,V., Ni,Z., Fedorova, M. Redox Biol.2015 Aug; 5: 195–204.
45. D.J.Betteridge, Metabolism, 2000, 49(2), 3–8.
46. B.Halliwell, Bio-chemical Journal, 2007, 401, 1.
47. Zheng J, Bizzozero OA. Journal of neuroscience research. 2010;88:3376–3385.
48. Mukherjee, M., Chio, TI., Sackett, DL., Bane, SL. Free Radic Biol Med. 2015 Jul; 84: 11–21.
49. D. Ray and P. K. Bharadwaj, Inorg. Chem.,2008, 47, 2252–2254.
50. S. L. Lin, P. Y. Kuo and D. Y. Yang, Molecules, 2007, 12, 1316–1324.
51. C. P. Kulatilleke, S. A. de Silva and Y. Eliav, Polyhedron, 2006, 25, 2593–2596.
52. Li, J., Zhang, CF .,Ming, ZZ., Yang, WC., Yang, GF. RSC Adv.,2013, 3, 26059-26065.
53. Zhang, Z. X., Guo, X. F., Wang, H. & Zhang, H. S. Anal Chem87, 3989–95 (2015).
54. Wu, M. Y. et al. Biomaterials 53, 669–78 (2015).
55. Feng, X., Liu, L., Wang, S. & Zhu, D. ChemSoc Rev 39, 2411–9 (2010).
56. Maruyama, S., Kikuchi, K., Hirano, T., Urano, Y. & Nagano, T..J Am ChemSoc124, 10650–1 (2002).
57. Woodroofe, C. C., Masalha, R., Barnes, K. R., Frederickson, C. J. & Lippard, S. JChemBiol11, 1659–66 (2004).
58. Wang et al., Scientific reports. 2016, 6, Article number: 23061.
59. Vegesna, G. K. et al.ACS Appl Mater Interfaces 2013,5, 4107–12.
60. Qin, W. et al.Advanced Functional Materials2012, 22, 771–779.
61. Boens, N., Leen, V. & Dehaen, W. Chem Soc Rev2012, 41, 1130–72.
62. Ziessel R, Ulrich G, Harriman A. New Journal of Chemistry, 2007, 31(4): 496–501.
63. Baruah M, Qin W, Vallée R A, Beljonne D, Rohand T, Dehaen W, Boens N. Organic Letters, 2005, 7(20): 4377–4380.
64. Bricks J L, Kovalchuk A, Trieflinger C, Nofz M, Büschel M, Tolmachev A I, Daub J, Rurack K. Journal of the American Chemical Society,2005, 127(39): 13522–13529.
65. Fan, G., Yang, L., Chen, Z. Frontiers of Chemical Science and Engineering,2014, Volume 8, 4, pp 405–417.
66. Dilek, O., Bane S. Bioorg Med Chem Lett.2009, 19(24): 6911–6913.
67. V. Ntziachristos, C. Bremer, R. Weissleder. Eur. Radiol.2003, 13, 195–208.
68. Y. Yang, Q. Zhao, W. Feng, F. Li. Chem. Rev.2013, 113 192–270.
69. Z. Guo, S. Park, J. Yoon, I. Shin. Chem. Soc. Rev. 2014, 43, 16–29.
70. Y.L. Pak, K.M.K. Swamy, J. Yoon. Sensors 2015, 15, 24374–24396.
71. K. Kiyose, H. Kojima, T. Nagano Chem. Asian J.2008, 3, 506–515.
72. Z. Xu, L. Xu. Chem. Commun.2016, 52, 1094–1119.
73. W. Sun, S. Guo, C. Hu, J. Fan, X. Peng. Chem. Rev. 2016, 116, 7768–7817.
74. Chin, J.,Kim HJ. Coordination Chemistry Reviews, 2017 (just published).
75. QCR Solutions Corp., http://www.qcrsolutions.com/Site/Near_Infrared_%28NIR%29_Dyes___QCR_Solutions_Corp.html
76. S. Atilgan, T. Ozdemir, E.U. Akkaya, Org.Lett. 2008, 10, 4065–4067.
77. Z. Dost, S. Atilgan, E.U. Akkaya, Tetrahedron 2006, 62, 8484–8488.
78. K. Umezawa, D. Citterio, K. Suzuki. Anal. Sci.2008, 24, 213–217.
79. I.A. Karpenko, M. Collot, L. Richert, C. Valencia, P. Villa, Y. Mély, M. Hibert, D. Bonnet, A.S. Klymchenko. J. Am. Chem. Soc. 2015, 137, 405–412.
80. Liu XD,Sun R., Ge GF.Xu YJ.,XuY., Lu JM.Org. Biomol. Chem., 2013,11, 4258-4264.
81. C. S. Yeh et al., Biomaterials. 2013, 34(22), 5677–5688.
82. J. V. Frangioni, Curr. Opin. Chem. Biol.2003, 7(5), 626–634.
83. J. Mieszawska et al., Bioconjugate Chem.2013, 24(9), 1429–1434.
84. A. Papadia et al., J. Minimally Invasive Gynecol.2015, 22(7), 1304–1306.
85. C.G. Williams. Trans. R. Soc. Edinburg, 1857, 21, 377–401.
86. X. Yi et al., Med. Sci. Monit.2015, 21, 511–517.
87. C. Shao et al., PloS One. 2014, 9(2), e88967.
88. G. Liberale et al., Eur. J. Surg. Oncol.2015, 41(9), 1256–1260.
89. A. K. Kirchherr, A. Briel, and K. Mader, Mol. Pharmaceutics. 2009, 6(2), 480–491.
90. O. A. Andreev et al., Proc. National Acad. Sci. U. S. A.2007, 104(19), 7893–7898.
91. J. A. Yang et al., Acta Biomater.2015, 12, 174–182.
92. S. Keereweer et al., J. Surg.Oncol. 2012, 105(7), 714–718.
93. M. Xu et al., Int. J. Cancer. 2012, 131(6), 1351–1359.
94. J. Gao et al., Biomaterials2011, 32(8), 2141–2148.
95. S. Luo et al., Biomaterials2011, 32(29), 7127–7138.
96. Shi C.,Wu JB., Pan D. Journal of Biomedical Optics2016, 21(5), 050901.
97. https://www.lumiprobe.com/tech/cyanine-dyes
98. Shimuzu et al., Nanomedicine: Nanotechnology, Biology, and Medicine, 2013.
99. Y. Yang, Q. Zhao, W. Feng, F. Li, Chem. Rev.2013113, 192–270.
100. X. Li, X. Gao, W. Shi, H. Ma, Chem. Rev. 2014, 114, 590–659.
101. X. Chen, F. Wang, J.Y. Hyun, T. Wei, J. Qiang, X. Ren, I. Shin, J. Yoon, Chem. Soc.Rev. 2016, 45, 2976–3016.
102. M. Vendrell, D. Zhai, J. Chen Er, Y.T. Chang, Chem. Rev. 2012, 112, 4391–4420.
103. L. You, D. Zha, E.V. Anslyn, Chem. Rev. 2015, 115, 7840–7892.
104. D. Wu, L. Chen, N. Kwon, J. Yoon, Chem 12016, 674–698.
105. S. Panda, A. Panda, S.S. Zade, Coord. Chem. Rev.2015, 300, 86–100.
106. L. Gai, J. Mack, H. Lu, T. Nyokong, Z. Li, N. Kobayashi, Z. Shen, Coord. Chem.Rev.2015, 285, 24–51.
107. V.B. Bregovic´ , N. Basaric´ , K. Mlinaric´-Majerski, Coord. Chem. Rev.2015, 295, 80–124.
108. D.-L. Ma, M. Wang, C. Liu, X. Miao, T.-S. Kang, C.-H. Leung, Coord. Chem. Rev.2016, 324, 90–105.
109. J. Chan, S.C. Dodani, C.J. Chang, Nat. Chem.2012, 4, 973–984.
110. E. Galbraith, T.D. James, Chem. Soc. Rev. 2010, 39, 3831–3842.
111. S. Banerjee, E.B. Veale, C.M. Phelan, S.A. Murphy, G.M. Tocci, L.J. Gillespie, D.O. Frimannsson, J.M. Kelly, T. Gunnlaugsson, Chem. Soc. Rev.2013, 42, 1601–1618.
112. P.A. Gale, E.N.W. Howe, X. Wu, Chem 12016, 351–422.
113. Y. Chen, R. Guan, C. Zhang, J. Huang, L. Ji, H. Chao, Coord. Chem. Rev.2016, 310, 16–40.
114. Wu D., Chen L., Lee W., KoG.,Yin J., Yoon J. Coordination Chemistry Reviews,2017
115. J. Yin, Y. Hu, J. Yoon, Chem. Soc. Rev.2015, 44, 4619–4644.
116. G.R.C. Hamilton, S.K. Sahoo, S. Kamila, N. Singh, N. Kaur, B.W. Hyland, J.F. Callan, Chem. Soc. Rev. 2015, 44 4415–4432.
117. T. Clausem, Physiol. Rev.2003, 83, 1269–1324.
118. Q. Yang, T. Liu, E.V. Kuklina, W.D. Flanders, Y. Hong, C. Gillespie, M.-H. Chang, M. Gwinn, N. Dowling, M.J. Khoury, F.B. Hu, Arch. Intern. Med.2011, 171, 1183–1191.
119. V.M. Vitvitsky, S.K. Garg, R.F. Keep, R.L. Albin, R. Banerjee, Biochim. Biophys. Acta 1822 (2012) 1671–1681.
120. H. Wulff, B.S. Zhorov, Chem. Rev.2008, 108, 1744–1773.
121. Y. Liu, Q. Su, M. Chen, Y. Dong, Y. Shi, W. Feng, Z.-Y. Wu, F. Li, Adv. Mater.2016, 28, 6625–6630.
122. Garland et al., Cell Chemical Biology, 2016.
123. Rodriguez et al., Trends in Biomedical Sciences,2017, 42, 2, 111-129. February 2017.
124. Kamiya M., Urano Y. Current Opinion in Chemical Biology2016, 33:9–15